09/09/19
Latin American subways ‘highest antimicrobial resistance’

By: Claudia Mazzeo
Send to a friend
The details you provide on this page will not be used to send unsolicited email, and will not be sold to a 3rd party. See privacy policy.
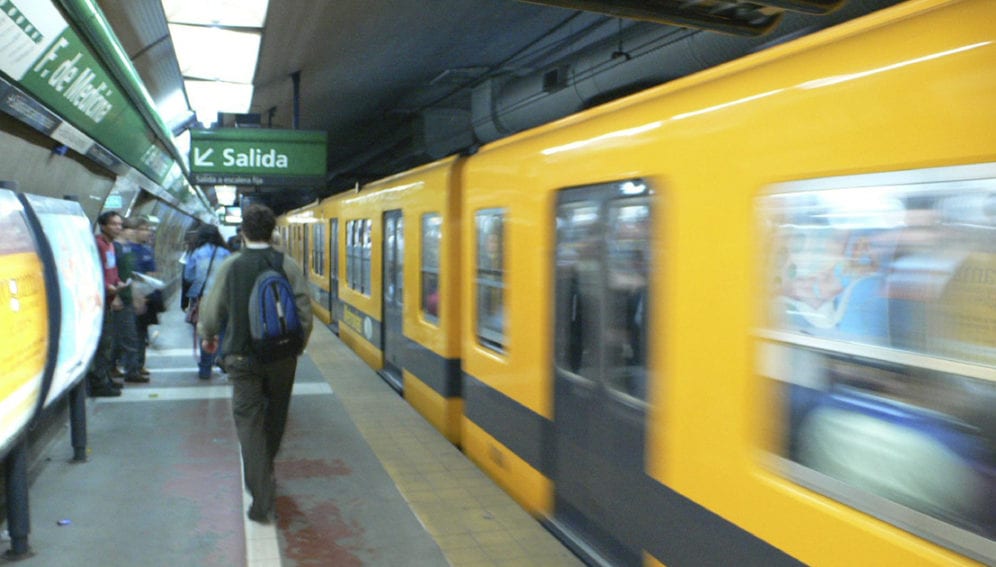
Tests taken on urban transport systems and subways in 58 cities around the world found Latin America to be the region with the greatest prevalence of antimicrobial resistance genes (RAM).
An international team of 600 researchers took 3,741 samples from the railings, ticket machines and walls of underground stations, to create an “atlas” of urban microorganism communities.
They used the samples to identify the bacteria, virus and fungi — known collectively as microbiota — in those urban places, and analyse variations in genetic characteristics and antibiotic resistance.
“This is an unprecedented study, which was possible thanks to technological advances that allow gene sequencing and computational analysis from a huge amount of data,”
Cristina Marino Buslje, scientist, Leloir Institute Foundation, Buenos Aires
Scientists identified 4,424 known species, of which 1,145 were found in more than 70 per cent of the samples. Sixty-one were found in more than 95 per cent of samples and are not part of the normal human microbiota from skin and airways, nor of the soil.
The study, by a consortium of scientists from Africa, America, Asia, Europe and Australasia, was published on BioRxiv, an open-access preprint repository that allows scientists to make their findings immediately available to the scientific community.
Eduardo Castro-Nallar, co-author of the paper and a researcher at Andrés Bello University’s Centre for Bioinformatics and Integrative Biology, in Santiago, Chile, said: “This led us to think that cities are an ecosystem by themselves, with a stable community of microorganisms.”.

Furthermore, the research found that more than 50 per cent of the collected genetic samples couldn’t be identified, so there are microorganisms not known or defined by science.
In Latin America, the scientists took samples from subways and urban transport systems of Sao Paulo, Rio de Janeiro, Bogotá, and Santiago. The prevalence of RAM genes found there was between 10 and 20 times higher than cities in other regions. Rio de Janeiro and Bogota, for example, showed ten times more RAM genes than Paris, Baltimore or Singapore.
The study also showed that the nature of RAM and the distribution of genes varies from city to city. “Though many microorganisms are present in many cities there is a sort of microbial signature that identifies every city: one could take a sample, analyse it and predict its origin,” said Castro-Nallar, who is also a researcher at George Washington University Computational Biology Institute.
“Some samples have just a few RAM genes, but others have a lot, which has consequences not only in how we understand urban development but also in urban planning,” he said.
This shows the need for better management of antibiotics in Latin America, the scientist said, adding: “The results of our paper indicate that the prevalence of RAM genes is very high, only surpassed by Offa, Nigeria.”
Cristina Marino Buslje, a scientist at the Leloir Institute Foundation’s Structural Bioinformatics Unit, in Buenos Aires, who did not participate in the study, said: “This is an unprecedented study, which was possible thanks to technological advances that allow gene sequencing and computational analysis from a huge amount of data.
“I hope that the data generated in this global study will be taken into account by government officials for decision-making in health policy. It could also help health professionals in the diagnosis and treatment of infections.”Virginia Pasquinelli, from the National Northwestern University of Buenos Aires Province, said: “This work not only brings new information but also provides free tools to urban microbiome analysis. The ultimate goal is to build a database accessible to the entire scientific community that allows replicating and validating information in different environments, generating benefits for public health.
“Nowadays, we know that microbiome modulates our immune system response; even how we respond to tumors is regulated by the diversity of our normal flora. To think that there is interaction with another urban microbiome is another good point to have in mind in further investigations.”