By: Carol Campbell
Send to a friend
The details you provide on this page will not be used to send unsolicited email, and will not be sold to a 3rd party. See privacy policy.
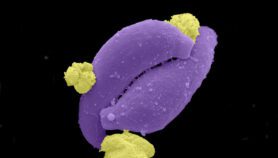
[OUDTSHOORN] A deadly type of Salmonella that is killing one in four infected people in Africa is a new strain, scientists have discovered.
Salmonella enterica Typhimurium is a foodborne bug that causes diarrhoea and is usually non-fatal. The strain found in Africa was thought to kill only people with compromised immune systems.
But now a collaboration between African and UK scientists to genetically sequence the strain — called ST313 — has shown that it has mutated to become resistant to many commonly used drugs.
The bacterium is thought to have emerged in the last decade and has since infected people in West, East and Central Africa. In 2002 it represented 95 per cent of S. Typhimurium found in Africa.
People develop fever and often arrive at clinics with blood poisoning. Even when treated with drugs to which it is sensitive, the bacterium kills one in four people.
The collaboration — between the Wellcome Trust Sanger Institute, the Malawi-Liverpool-Wellcome Trust Clinical Research Programme, which first found the bacteria, and the KEMRI-Wellcome Trust Programme — analysed 50 samples of bacterial DNA from the blood of patients with severe symptoms of infection and who were also suffering from HIV, malaria, malnutrition or anaemia.
They compared the genetic sequence with those of strains associated with milder disease and found that ST313 had acquired a set of genes that makes it resistant to many antibiotics.
Robert Heyderman, director of the Malawi-Liverpool-Wellcome Trust programme, told SciDev.Net that the genetic sequence suggested the bacteria could be passed from human to human.
"Salmonella is a bacteria which we know is passed on to humans from eggs, meat or milk, but we think this bacteria could be coming off cutlery and crockery. It’s in the home and everyone, especially small children, is vulnerable," he said.
Heyderman added that genetic sequencing is important because only by understanding the cause of diseases can health authorities take action to protect people. But surveillance for such diseases is poor outside South Africa, so no one knows how many people have been infected or died.
Anthony Smith, a gut disease scientist at South Africa’s National Institute for Communicable Diseases, a division of the National Health Laboratory Service, said work is underway to establish an international database of Salmonella DNA fingerprints.
"There is a dearth of information from Africa," he added. To fingerprint a Salmonella sample, "you need molecular laboratories, apparatus and scientists which don’t exist in many other countries on the continent".