Send to a friend
The details you provide on this page will not be used to send unsolicited email, and will not be sold to a 3rd party. See privacy policy.
Several promising drug development strategies are emerging in the fight against tuberculosis (TB), which continues to claim nearly two million lives worldwide each year, largely in developing countries.
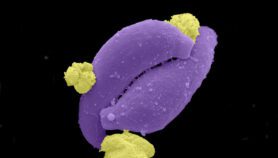
As the bacterium — Mycobacterium tuberculosis (Mtb) — evolves to resist anti-TB drugs, and patients find it difficult to stick to existing drug regimes the search is on for both new drugs and good diagnostics.
Some promising drugs are in early clinical trials — for example PA-824, which could reduce the time needed to treat TB — but the odds are stacked against them because fewer than ten per cent of antibiotics that enter early clinical trials ever gain approval.
But researchers are finding that "omics" — fields involving the study of genes, proteins and metabolism — offer the potential to unlock new information about the bacterium and its interactions with people in unprecedented detail.
As well as genomics, proteomics and metabolomics, scientists are also using chemical genomics, an approach that reverses the standard drug discovery process.
Mtb lives among plenty of competitor bugs equipped with their own weapons to keep the bug in check. Scientists are trying to understand how these rival bacteria have evolved and are applying modern omics tools to identify the defensive molecules, screen them for their anti-TB potential and pinpoint just where they strike at Mtb.
This approach "could well uncover entirely new classes of drugs".
Other fields seek to enhance knowledge about the bacterium itself. Structural genomics, for example, aims to uncover the three-dimensional structure of every protein in Mtb.
The ultimate goal is to replicate Mtb "in silico"— that is, "produce a computer simulation of the bacterium that behaves just like the real thing does in the body". Scientists should then be able to predict which bacterial components will make the best drug targets — and which candidate drugs will hit those targets most effectively.
Researchers are confident that, based on current progress,they will be able to achieve this within the next 20 years.