By: Baraka Rateng’
Send to a friend
The details you provide on this page will not be used to send unsolicited email, and will not be sold to a 3rd party. See privacy policy.
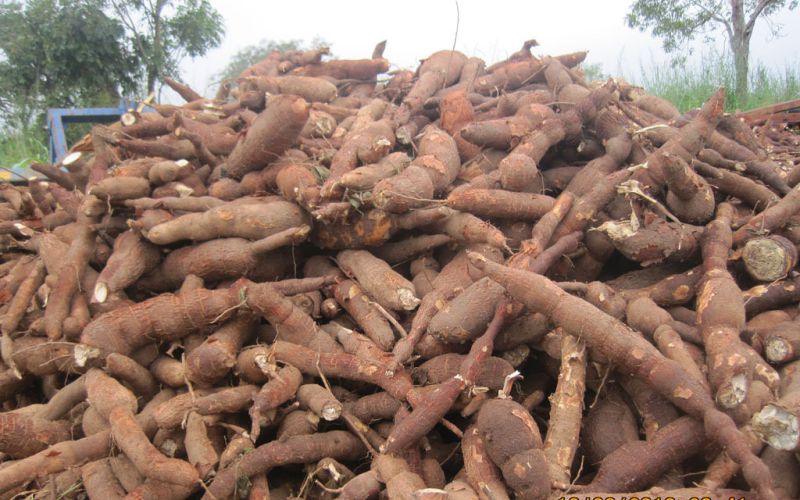
Researchers have sequenced the genome of the cassava — enabling them to better understand the genetic basis the plant’s disease resistance, quality and crop maturity.
A research team in Kenya spent four years decoding the DNA, the genetic material, of the cassava plant, which is widely farmed and eaten across the tropics.
“Cassava is the main food security crop of [Africa], so providing a high yield in poor soils with minimal water can be crucial when other crops fail.”
Morag Ferguson, IITA
The researchers identified the order of the genetic letters of 53 cultivated and wild cassava plant materials from Africa, Asia, South America and Oceania — a process called genome sequencing. They also sequenced five cassava-related plants such as M. glaziovii and identified the genetic components of 268 African cassava varieties.
The study which started in 2012, was aimed at increasing the genomic resources for cassava, says Morag Ferguson, a co-author of the study and a molecular geneticist at the International Institute of Tropical Agriculture (IITA), Kenya. The research was published in the journal Nature Biotechnology last month
“The study includes 97 per cent of the estimated genes,” Ferguson tells SciDev.Net. “The large amount of DNA sequence information provides insights into the origin of cassava and resources for the improvement of cassava”.
For example, the genome holds information on resistances to cassava brown streak disease, a devastating viral disease affecting cassava in southern, eastern and central Africa.
Sequence information, according to Ferguson, revealed that some disease-resistant cassava varieties in Tanzania, including Namikonga and Muzege, contain sections of genomes of M. glaziovii.
“Cassava is the main food security crop of [Africa]”, so providing a high yield in poor soils with minimal water can be crucial when other crops fail, says Ferguson.
The study was conducted by researchers from countries as varied as Fiji, Kenya, Micronesia, Nigeria, Tanzania and United States.
Paul Kimani, a plant breeder from Kenya’s University of Nairobi says the main contribution of the findings is a clear demonstration of the genetic relationships among the various species, including cultivated cassava, its wild relatives and others in the secondary or even tertiary gene pool.
“What it does not do is link the genes with any economically important traits such as disease resistance, nutritional quality or agronomic traits,” he says.
Kimani explains that because cassava breeders often have little genetic variety among their crop, an epidemic can easily cause severe damage, leading to rapid spread of diseases such as cassava mosaic disease and brown streak disease in Africa.
This piece was produced by SciDev.Net’s Sub-Saharan Africa English desk.
References
Jessen V. Bredeson and others Sequencing wild and cultivated cassava and related species reveals extensive interspecific hybridization and genetic diversity (Nature Biotechnolgy, 18 April 2016)