Send to a friend
The details you provide on this page will not be used to send unsolicited email, and will not be sold to a 3rd party. See privacy policy.
Studying bacteria hosted by wildlife helps to detect and control the spread of antimicrobial resistance across ecosystems and back to humans, they say.
As each species interacts with its environment in different ways, scientists examine these patterns to identify where antibiotic exposure occurs and how humans contribute to this process through their sanitation habits, says Kathleen Alexander, a disease ecologist at Virginia Tech in the United States.
Alexander and her colleague Sarah Jobbins studied E. coli — a bacterium commonly found in the foods and intestines of people and animals — to gauge the spread of antibiotic resistance among humans, domestic animals and wildlife in Chobe National Park and two villages in northern Botswana.
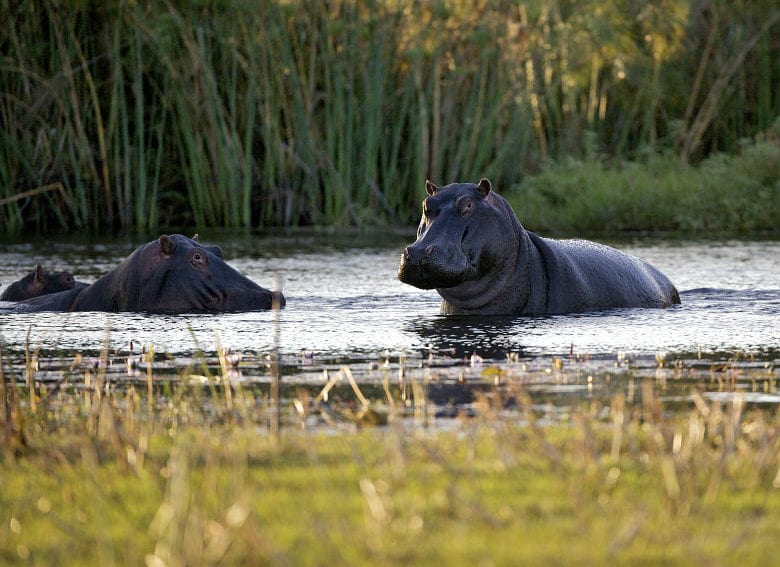
“Water seems to be the most important medium exposing animals to antibiotic resistance, with water-associated species such as hippos and waterbucks having higher levels of multi-drug resistant microorganisms.”
Kathleen Alexander, Virginia Tech
Forty-one per cent of the faecal samples from 18 wild species contained E. coli resistant to one or two of ten antibiotics tested, says the study, published in the Journal of Wildlife Diseases on 31 July.
Samples from domestic cattle, healthy and ill people, and environmental sources of human faeces such as bush latrines and sewage sludge were also tested. No antimicrobial resistance was detected in the cattle faeces. But the researchers found that bacteria from wildlife, humans and the environmental samples were resistant to a similar range of antibiotics.
“Resistance was widespread but not ubiquitous among sampled animals,” the paper says.
“Water seems to be the most important medium exposing animals to antibiotic resistance, with water-associated species such as hippos and waterbucks having higher levels of multi-drug resistant microorganisms,” Alexander says.
In addition, bacteria resistant to multiple drugs were more common in animals, such as baboons, warthogs and mongooses, which live in urbanised areas, and in carnivorous species.
“Resistance may accumulate up the food chain, “making apex predators such as crocodile, leopard and hyena important ecosystem sentinels”, Jobbins said in a statement last month.
Studying wild animals helps to understand how resistance moves from humans and farming systems across ecosystems and ultimately back to humans, says Alexander, who also co-heads the Center for African Resource: Animals, Communities and Land Use, a non-profit conservation agency in Botswana.
According to the paper, this approach could be applied to other ecosystems to improve the early detection of antimicrobial resistance epidemics.
Studying antibiotic resistance in humans, livestock and the environment at once helps to better understand the links between the three, says Grace. She adds that it would be useful to repeat similar research to observe trends over several months or years.
References
Sarah Elizabeth Jobbins and Kathleen Ann Alexander Whence they came — antibiotic-resistant Escherichia coli in African wildlife (Journal of Wildlife Diseases