By: Esther Nakkazi
Send to a friend
The details you provide on this page will not be used to send unsolicited email, and will not be sold to a 3rd party. See privacy policy.
[KAMPALA] A study has suggested that scientists could predict where on the planet the next virus could jump from animals to humans, thus providing data that will help in early warning systems and disease surveillance efforts.
According to the US-based researchers, few analytical tools exist to help scientists understand the patterns of viral diversity in wildlife and how these may successfully become the next human virus, or which viruses could cross species boundaries.
The study, published last month (21 June) in the journal Nature, helps build a roadmap of where to prioritise disease surveillance efforts around the world to better stop viruses from having a large impact.
In the study, the scientists have mapped out the ‘missing zoonoses’ giving geographic hotspots as eastern, central and southern Africa, South and Central America as well as parts of Asia.
According to the study, ‘missing zoonoses’ are those viruses that could jump from animals to humans which were previously unknown. The map it produced shows locations likely to be the source of the next emerging zoonotic disease.
“We want to get ahead of the curve, and be able to develop early warning systems to identify and find emerging disease threats.”
Kevin J. Olival
The authors analysed major online databases such as PubMed and Google Scholar for articles published between 1940 and 2015 for zoonotic viruses — those known to have been detected in humans and at least one other mammalian host. They also reviewed books, reviews, and literature cited in sources of the identified articles.
They then created a database of 2,805 mammal–virus associations to build mathematical models that allowed them to identify which species traits and groups of mammals carry the largest numbers of viruses and the virus traits that make some viruses more likely to jump into humans.
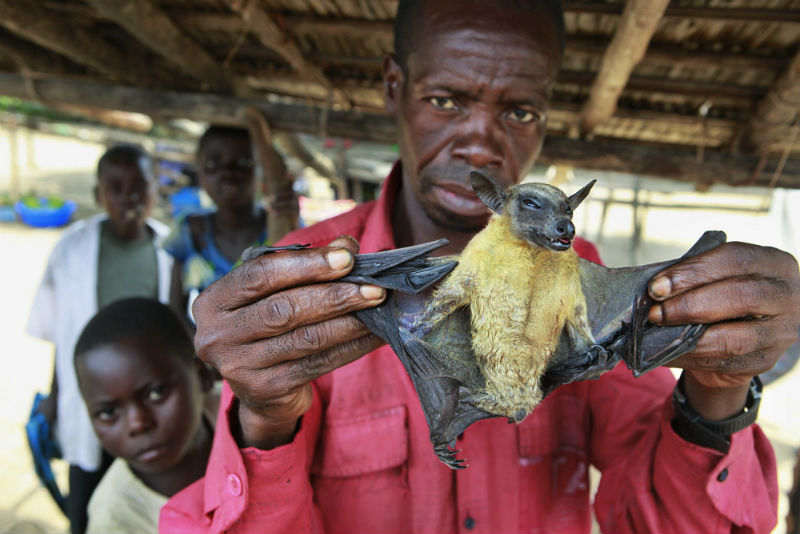
For instance, they demonstrated that bats harbour a significantly higher proportion of zoonotic viruses than all other mammals.
“We want to get ahead of the curve, and be able to develop early warning systems to identify and find emerging disease threats before or soon after they emerge in the human population,” says Kevin J. Olival, the principal investigator and an associate vice-president for research for US-based EcoHealth Alliance, which conducted the study.
“These show which are the predicted number of viruses that are out there, minus what we already know exist. We also identify virus traits that make some viruses more likely to jump into humans versus others.”
Olival adds that their data is already in use as part of a project to find and characterise new viruses around the world, and to better understand the location-specific risk factors and human behaviours in zoonotic virus hotspots.
Ritah Nakayinga, a lecturer in virology at the International Health Sciences University in Uganda, tells SciDev.Net that the study showed that classes of mammals such as primates and rodents have a higher proportion of zoonotic virus, which is an interesting finding and is supported by the Ebola and Marburg outbreaks in the world and Uganda respectively.
“Although much of the study focused on reservoir mammals, exploration into insect vectors would throw light on spillover events which are the basis of recent outbreaks [such as] Zika, Chikugunya and yellow fever,” Nakiganda explains.
“This raises questions on methodologies used to conclude this,” says Nakayinga. “It goes to show that literature-based databases for establishment of models may not be appropriate tools for making predictions although they may be preferred because of costs implications.”
This piece was produced by SciDev.Net’s Sub-Saharan Africa English desk.
References
Kevin J. Olival and others Host and viral traits predict zoonotic spillover from mammals (Nature, 21 June 2017)